-

Adding citations between existing articles in Wikidata
Scholarly articles provide context to the factualness of statements in Wikidata, similar to the [citation needed] in Wikipedia. And just like the cited references in each scholarly article itself. The citation network is general seen as an essential part of (doing) science, even without citation intention annotation. Nowadays, citations are mostly open, but this took very serious lobbying by the Initiative for Open Citations and not every publisher reacted immediately. But now that they are open, projects like OpenCitations are making this citation network FAIR. -
Scholia configurability
Scholia is a visual layer on top of Wikidata providing a rich user experience for browing scholarly research related knowledge. I am using the combinatie for various things, including exploring new research topics (a method, compound, or protein I do not know so much about yet), indexing notable research output (including citations), progress of Citation Typing Ontology uptake, etc. This weekend I hope to send around the final draft for the Scholia Chemistry paper. -
Kasabi archive at the Internet Archive
Kasabi was an innovative RDF publishing platform from around 2011. Shortlived, and maybe just too early. I published two open datasets there. One was ChEMBL-RDF (see these posts). The second was a small data sets called ChemPedia, a open science effort to crowdsource chemical names. This is still very much needed, and possibly Wikidata could fill that gap, but it would first need to be able to handle all labels as statements itself. -
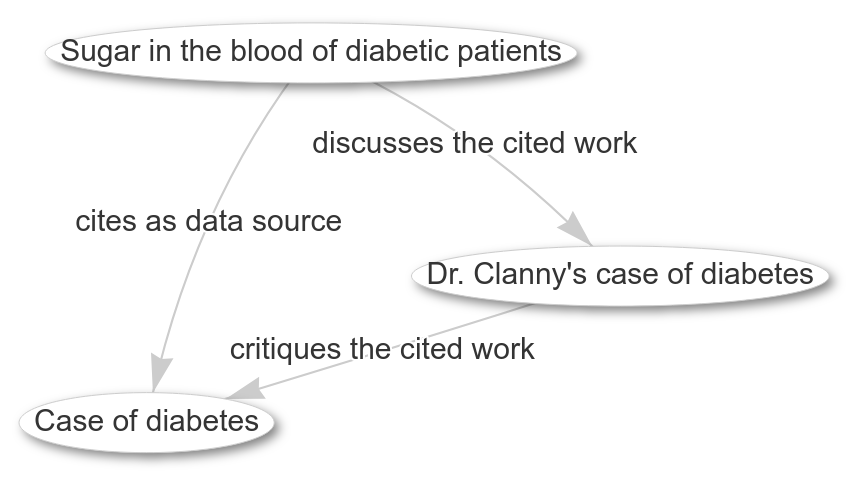
Scholarly discussions through the eyes of CiTO (and Wikidata)
Diabetes was already discussed in literature back in 1838-1839 (doi:10.1016/S0140-6736(02)96038-1, doi:10.1016/S0140-6736(02)96066-6, and doi:10.1016/S0140-6736(02)83966-6). These three papers show a short discussion. Papers were a lot shorter back in the days, and the discussion actually shows why papers are longer now (tho I am not sure they really got sufficiently more reproducible, but that’s another discussion). -
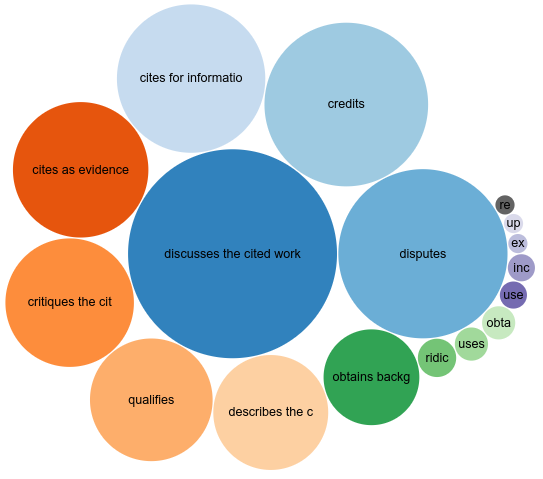
CiTO updates: Wakefield and WikiPathways
This summer I am trying to finish up some smaller projects that I did not have time for to finish, with mixed successes. I am combing this with a nice Dutch staycation, and I already cycled in Overijssel and in south-west Friesland and learning about their histories. But this post is about an update on my Citation Typing Ontology use cases. And I have to say, a mention by Silvio Peroni is pretty awesome, thanks! -
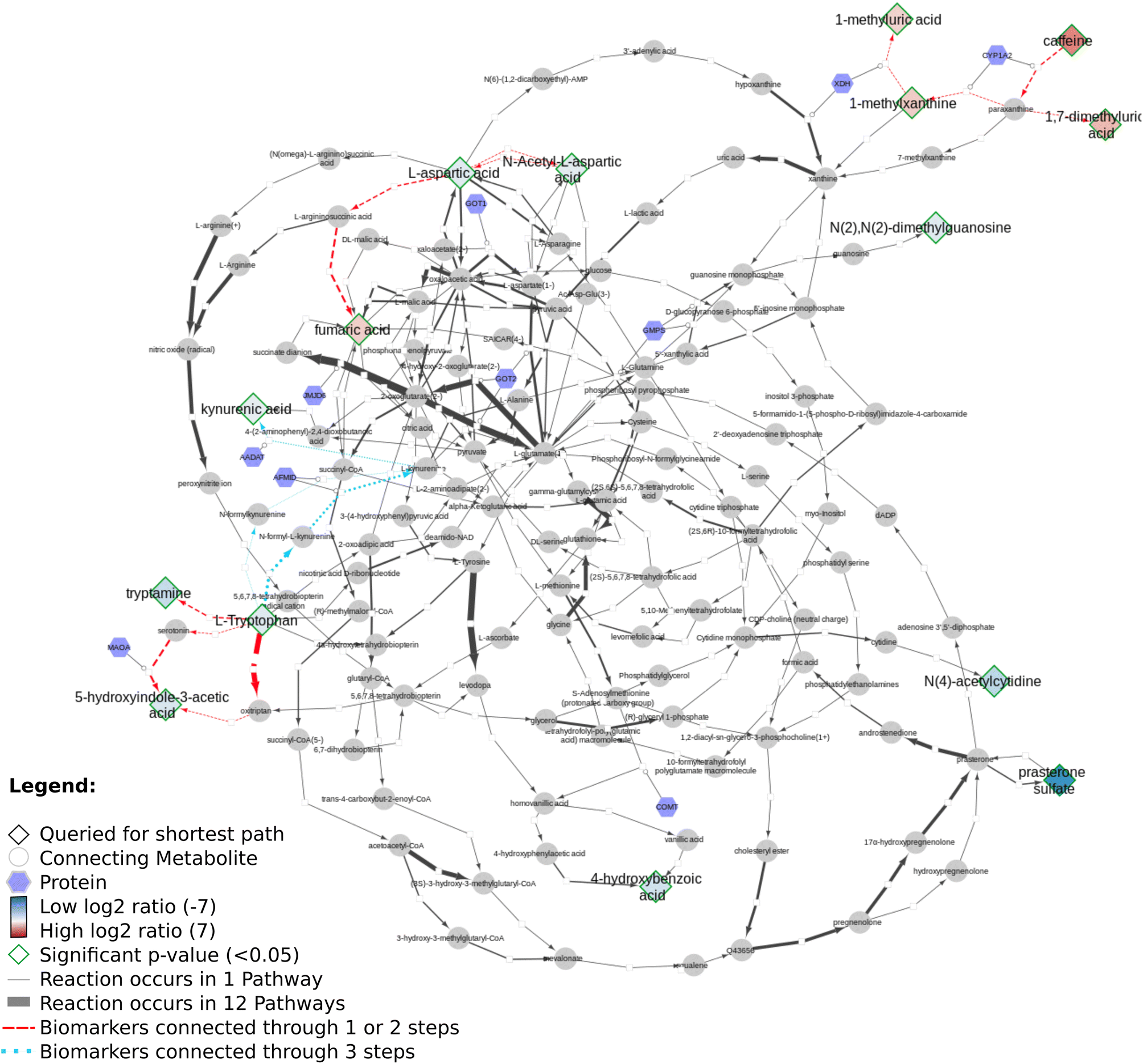
New paper: "Discovering life's directed metabolic (sub)paths to interpret human biochemical markers using the DSMN tool"
I am still catching up with a lot of work, and found out I actually had forgotten to blog about this cool article by Denise Slenter: “Discovering life’s directed metabolic (sub)paths to interpret human biochemical markers using the DSMN tool” (doi:10.1039/D3DD00069A). This paper explains how various open science resources (Wikidata, Reactome, WikiPathways) are used to visualize the biological story of the data from two metabolomics experiments archived in MetaboLights. -

GoatCounter, Rogue Scholar and more new things
About a year ago I started migrating my blogger.com blog to a git-version-controlled, Markdown-based blogging platform. I have to say, it has been a happy year. It actually is awesome to port old blog posts (follow that here) and to see what I have been working on some 17, 18 years ago.