Semantic Web features in Bioclipse 2.2
Ola is releasing Bioclipse 2.2.0 today, and asked me to show case the semantic web functionality in Bioclipse. I realized that I do not have a nice page showing the semantic web overview. But I did blog a lot about RDF functionality, so here’s a list of pointers:
- Bioclipse Manager for MyExperiment.org
- Bioclipse, RDF and defeasible reasoning (see also Samuel’s blog)
- Bioclipse and SPARQL end points #2: MyExperiment
- Bioclipse and SPARQL end points
- Solubility Data in Bioclipse #2: handling RDF
- Solubility Data in Bioclipse #3: Finding ChEBI IDs
- Solubility Data in Bioclipse #4: Finding ChEBI IDs (Again, but better)
- /me is having Bioclipse/XMPP/RDF fun
Or check this screenshot from a Posterous post about a MyExperiment workflow :
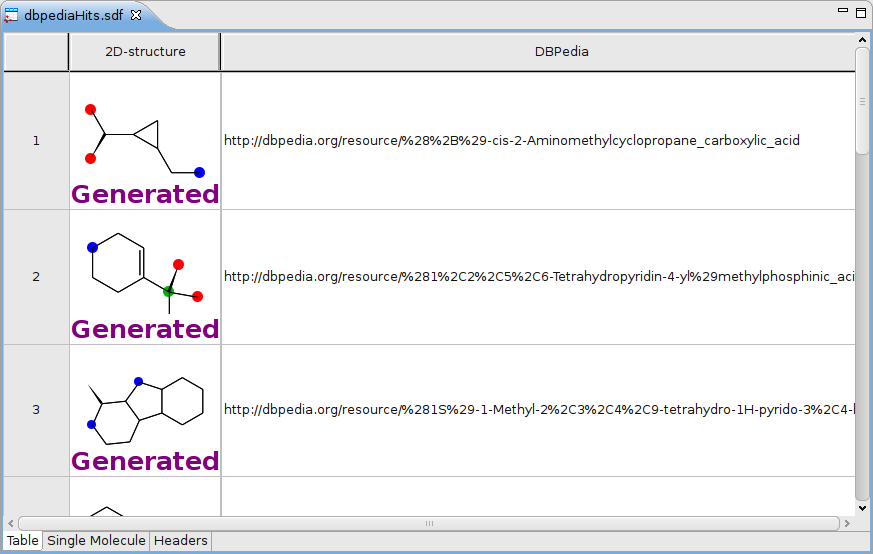
One thing I have not blogged about yet (I think), is that the Bioclipse RDF manager also understands RDFa now. Well, sort of… it relies on a webservice, but this is what the script looks like:
model = rdf.createStore()
rdf.importRDFa(model, "http://egonw.github.com/")
rdf.saveRDFN3(model, "/Virtual/egonw.n3")
With support of SPARQL end points, and reading RDF from web resources directly (RDF/XML, N3, RDFa), Bioclipse is ready for the chemical semantic web.