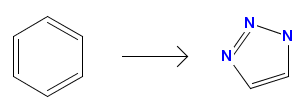
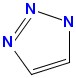
-
Oxford... #2
The Predictive Toxicology meeting is over. It was a great meeting, by any standard. Very much recommended, and many thanx to Barry for the organization! The meeting was a true workshop, with a mix of presentations and getting work done. I participated in a group that looked at mutagenicity of potential anti-malaria drugs from the datasets of GSK and Novartis recently release as Open Data. We used various tools to predict properties, and plan to make all our results freely available soon. Otherwise, it was also great to meet Nina again (with whom I talked about OpenTox), and to meet other CDK users, including Patrik (SMARTCyp , doi:10.1021/ml100016x) and David (Inkspot). -
Oxford...
Yesterday I arrived in Oxford, after a 3.5 hour bus transfer from London Stansted. Long, boring ride (though I might have seen a few red kites , but seeing that they were near extinct, I am wondering what other large bird of prey has strong split tail like a swallow). Showed once more that the UK infrastructure has hardly changed since the 19th century. Enjoying an undergraduate room at one of the colleges. Pretty basic, but makes me feel more like a human than a tourist. Yes!, undergraduate students are human too! One of the advantages is you get an excellent internet connection :) -
Cb: New Blogs #13
The Cb software is still holding… I jettinsoned the old post cache, which speeded up the processing of blogs considerably, but the system just doesn’t scale right. Yet, Euan has done a great job, and the Cb site has now been online for some three years! Here are some new blogs included in the aggregation and analysis: -
CDK-JChemPaint #3: rendering parameters
OK, one last CDK-JChemPaint tutorial for today (see #1 and #2). Rendering wasn’t as much fun, if you could not tune it to your needs. JChemPaint has long had many rendering parameters, and one by one we are converting them to the new API. The following code is an modification to the first example, and adds some code to list all rendering parameters for the three used generators: -
New Blue Obelisk Exchange online at Shapado.com
StackOverflow has served as very well in the past couple of months with the Blue Obelisk Exchange. The BOx was taking advantage of a beta project of StackOverflow, and users of that program can switch to a payed plan after the beta phase was over. That moment is nearing, but the pricing model is just unrealistic for us. There were also some comments on the Blue Obelisk using proprietary software (see Stackoverflow not open source — not a problem?).