-
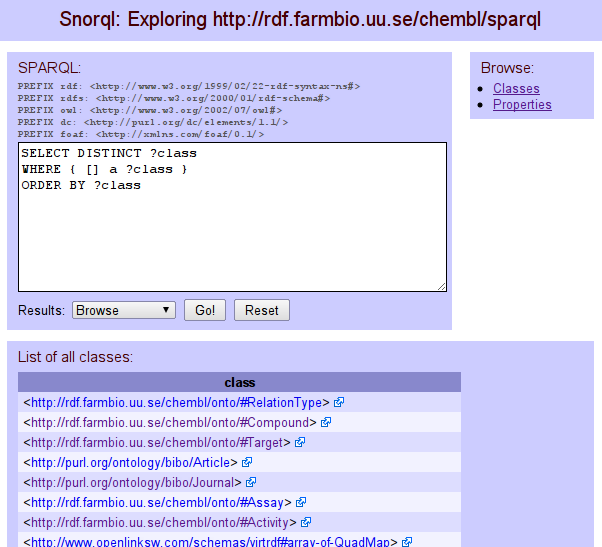
ChEMBL RDF #1:SPARQL end point
In a series of SPARQL end points , I am happy to present a new Virtuoso 6.1-hosted SPARQL end point for the ChEMBL database (CC-BY-SA), at our groups new rdf.farmbio.uu.se server. The server is hosting 23.8M triples, with the data based on ChEMBL 02. There is a SPARQL end point, as well as a SNORQL interface: -
UU Cheminformatics Journal Club
Following the steps of the IU Cheminformatics Journal Club, I have started a UU Cheminformatics Journal Club: