Data Curation: 5% inspiration, 95% frustration (cleaning up data inconsistencies)
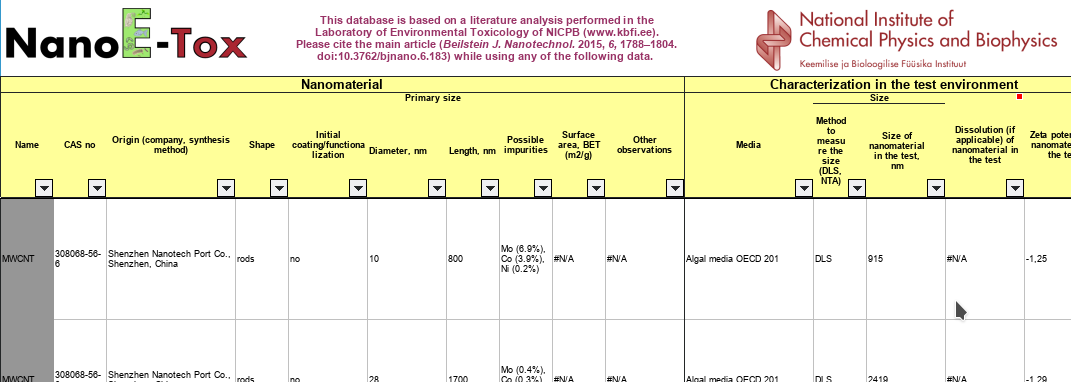
Slice of the spreadsheet in the supplementary info.
Just some bit of cleaning I scripted today for a number of toxicology end points in a database published some time ago the zero-APC Open Access (CC_BY) journal Beilstein of Journal of Nanotechnology, NanoE-Tox (doi:10.3762/bjnano.6.183).
The curation I am doing is to redistribute the data in the eNanoMapper database (see doi:10.3762/bjnano.6.165) and thus with ontology annotation (see doi:10.1186/s13326-015-0005-5):
recognizedToxicities = [
"EC10": "http://www.bioassayontology.org/bao#BAO_0001263",
"EC20": "http://www.bioassayontology.org/bao#BAO_0001235",
"EC25": "http://www.bioassayontology.org/bao#BAO_0001264",
"EC30": "http://www.bioassayontology.org/bao#BAO_0000599",
"EC50": "http://www.bioassayontology.org/bao#BAO_0000188",
"EC80": "http://purl.enanomapper.org/onto/ENM_0000053",
"EC90": "http://www.bioassayontology.org/bao#BAO_0001237",
"IC50": "http://www.bioassayontology.org/bao#BAO_0000190",
"LC50": "http://www.bioassayontology.org/bao#BAO_0002145",
"MIC": "http://www.bioassayontology.org/bao#BAO_0002146",
"NOEC": "http://purl.enanomapper.org/onto/ENM_0000060",
"NOEL": "http://purl.enanomapper.org/onto/ENM_0000056"
]
With 402(!) variants left. Many do not have an ontology term yet, and I filed a feature request.
Units:
recognizedUnits = [
"g/L": "g/L",
"g/l": "g/l",
"mg/L": "mg/L",
"mg/ml": "mg/ml",
"mg/mL": "mg/mL",
"µg/L of food": "µg/L",
"µg/L": "µg/L",
"µg/mL": "µg/mL",
"mg Ag/L": "mg/L",
"mg Cu/L": "mg/L",
"mg Zn/L": "mg/L",
"µg dissolved Cu/L": "µg/L",
"µg dissolved Zn/L": "µg/L",
"µg Ag/L": "µg/L",
"fmol/L": "fmol/L",
"mmol/g": "mmol/g",
"nmol/g fresh weight": "nmol/g",
"µg Cu/g": "µg/g",
"mg Ag/kg": "mg/kg",
"mg Zn/kg": "mg/kg",
"mg Zn/kg d.w.": "mg/kg",
"mg/kg of dry feed": "mg/kg",
"mg/kg": "mg/kg",
"g/kg": "g/kg",
"µg/g dry weight sediment": "µg/g",
"µg/g": "µg/g"
]
Oh, and don’t get me started on actual values, with endpoint values, as ranges, errors, etc. That variety is not the problem, but the lack of FAIR-ness makes the whole really hard to process. I now have something like:
prop = prop.replace(",", ".")
if (prop.substring(1).contains("-")) {
rdf.addTypedDataProperty(
store, endpointIRI, "${oboNS}STATO_0000035",
prop, "${xsdNS}string"
)
rdf.addDataProperty(
store, endpointIRI, "${ssoNS}has-unit", units
)
} else if (prop.contains("±")) {
rdf.addTypedDataProperty(
store, endpointIRI, "${oboNS}STATO_0000035",
prop, "${xsdNS}string"
)
rdf.addDataProperty(
store, endpointIRI, "${ssoNS}has-unit", units
)
} else if (prop.contains("<")) {
} else {
rdf.addTypedDataProperty(
store, endpointIRI, "${ssoNS}has-value", prop,
"${xsdNS}double"
)
rdf.addDataProperty(
store, endpointIRI, "${ssoNS}has-unit", units
)
}
But let me make clear: I can actually do this, add more data to the eNanoMapper database (with Nina), only because the developers of this database made their data available under an Open license (CC-BY, to be precise), allowing me to reuse, modify (change format), and redistribute it. Thanks to the authors. Data curation is expensive, whether I do it, or if the authors of the database did. They already did a lot of data curation. But only because of Open licenses, we only have to do this once.