Uncertainty in NMR based 3D protein models
While I was working on implementing proper author-given chain IDs in PDB structures for Jmol’s mmCIF reader today, I thought it was interesting to mention the recent article Traditional Biomolecular Structure Determination by NMR Spectroscopy Allows for Major Errors by Nabuurs (DOI:10.1371/journal.pcbi.0020009, open access), working at the CMBI, two floors away from my former working location at the Radboud University Nijmegen.
Nabuurs discusses in this article the uncertainties that come with NMR derived 3D molecular structures of proteins. These studies do not give factual data on atomic coordinates, but generally give facts about interatomic distances. Solving the 3D geometry is then an optimization problem where the task is to find the 3D geometry that best reproduces the factual interatomic distances.
Now, this optimization has many closeby, i.e. in terms of matching the experimental data, minima, corresponding, possibly, to quite different structures.
This is nicely demonstrated in the article, by comparing the folds of 1Y4O and 1TGQ, as shown in the figure below (CCAL license):
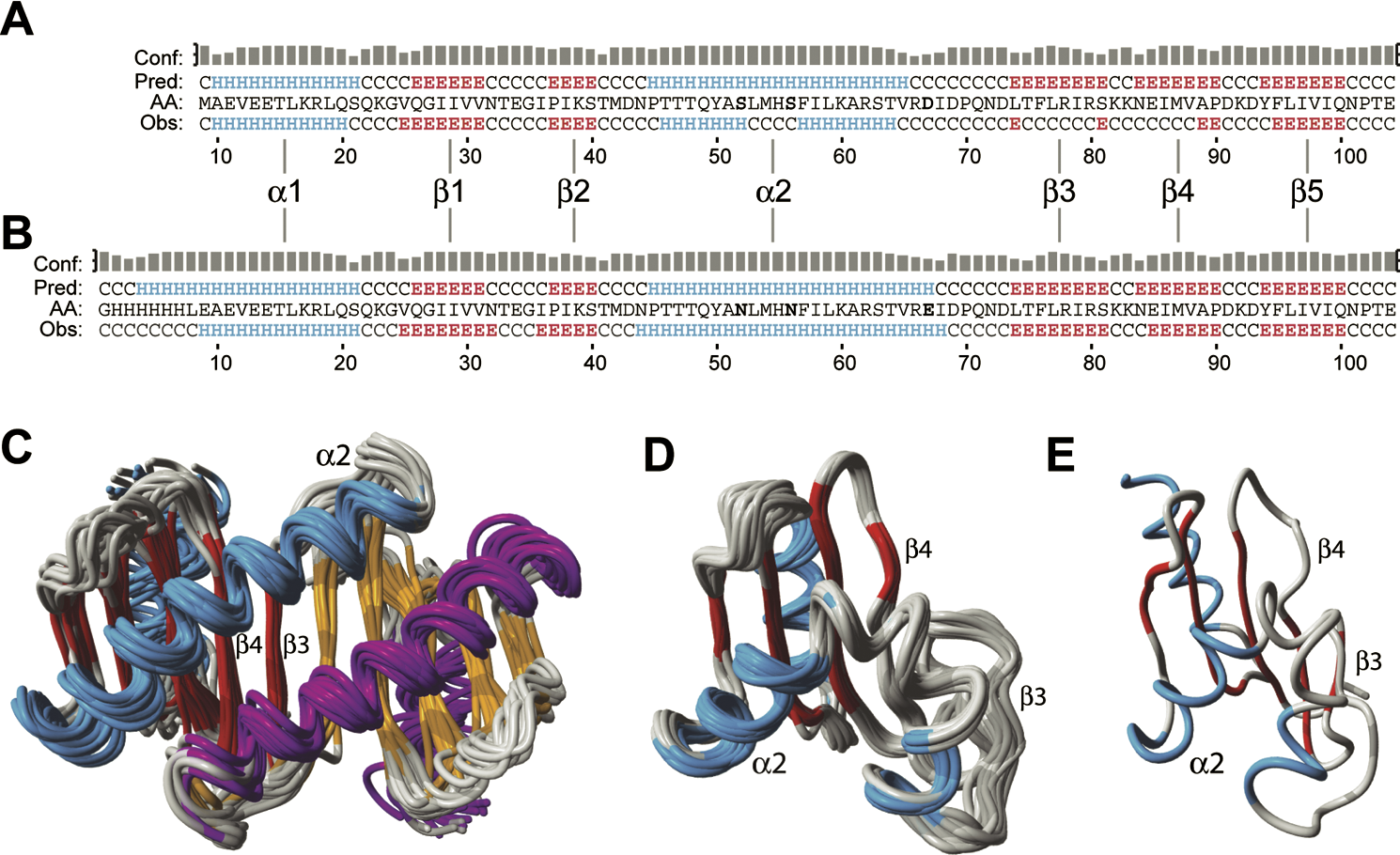
It is interesting to note that 1TGQ got replaced by 2B95 about the same time the article by Nabuurs was published, which shows a 3D model that is homologous with that of 1Y4O, and different from that in the Nabuurs article.