In a comment, Henry Rzepa proposed inclusion of CML, and refers to earlier work on CMLRSS where Chemical Markup Language is embedded in RSS news feeds for which I wrote readers for Jmol and JChemPaint (DOI:10.1021/ci034244p).
As readers of my blog know, the Bioclipse project has been working hard on an integrated (bio)chemistry workbench, and the latest release includes a CMLRSS reader plugin too, which supports CML embedded in Atom 0.3/1.0 and RSS 1.0/2.0 feeds. Now, adding support for other embedded namespaces is trivial, and this morning I hacked in support for OTMI:
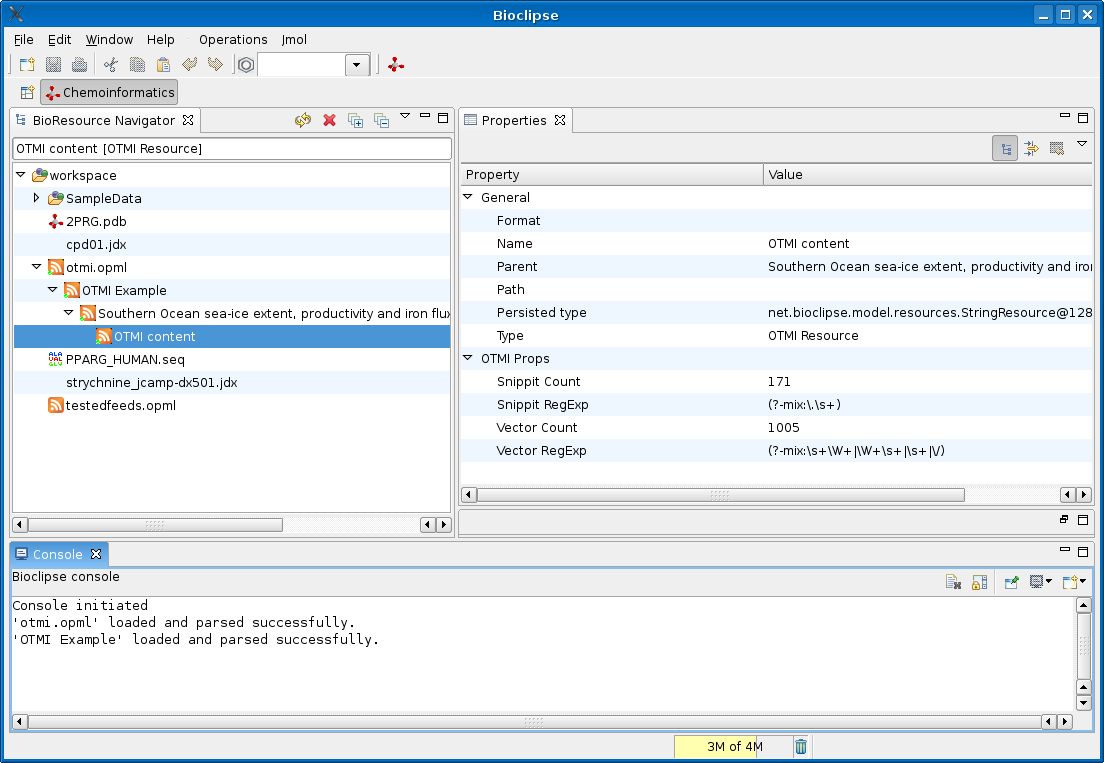
This screenshot show the original OTMI example
with the Atom 1.0 entry now wrapped in an Atom 1.0 <feed> element. There is no nice OTMI icon for the OTMI content in the
Atom 1.0 entry, neither did I make a ‘view’ yet showing the actual vector’s or the snippet’s, but that’s a piece of cake too.
Now, the nice thing about this is that the Bioclipse code for the Atom and RSS feeds, just greps through the feed entry and show whatever CML or OTMI content is present. When Nature decides to include CML in these OTMI files too, I will not have to update the current code.
]]>