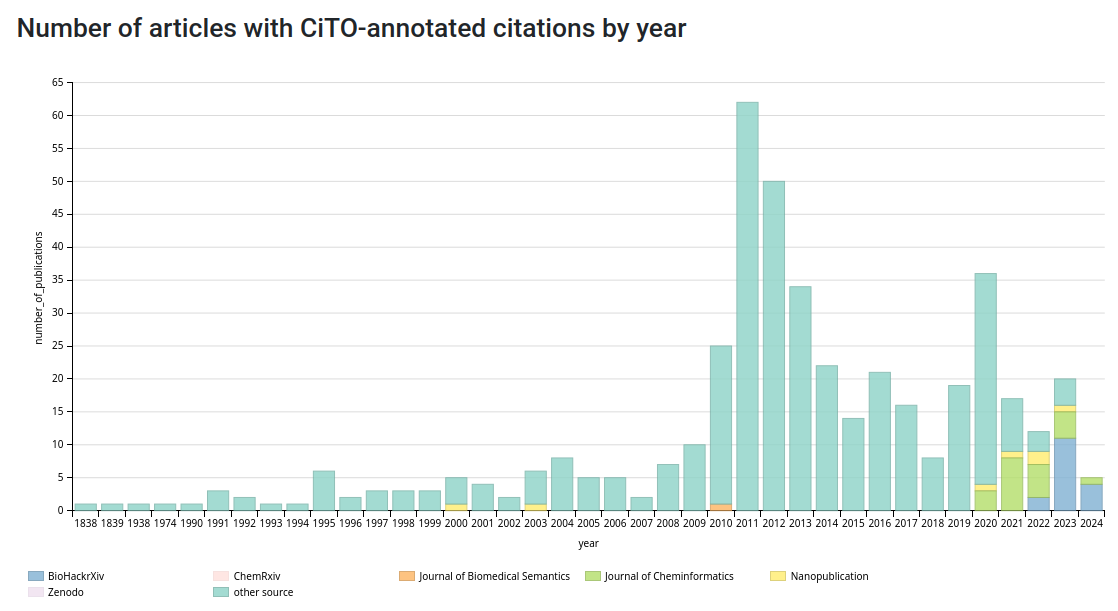
Now, in this post I will document an update of how we can use nanopublications for citation intention annotation, and compare this to existing solutions. I have been collecting and indexing the CiTO intention annotations in Wikidata and visualizing the corpus with Scholia at scholia.toolforge.org/cito/. There are currently 22 journal articles with explicit CiTO annoation, largely thanks to a Journal of Cheminformatics pilot (e.g. see doi:10.1186/s13321-023-00683-2). Recently, the preprint/report server BioHackrXiv started CiTO support too, also visible in the statistics on Scholia with another 17 papers. A third source is data sets from bibliometric-like studies, as explained in this post. Nanopublications would be a fourth solution.
So, why another solutions? Like the datasets, assuming DataCite approaches, have clear provenance, but the overhead of and needed time for creating a dataset with citation intent annotations can be limiting. And because nanopublications can be linked to ORCID identifiers, we can even discover which citation intent annotations are created by the original authors of articles. Another advantage is that nanopubs are basically RDF and we can query them easily, allowing the citation intentions to migrate to Wikidata. Scholia already saw an update to recognize nanopublications as a unique kind reference (see the new Wikidata property Nanopublication identifier (P12545)).
NanoDash template
So, if we can make it easy for people to define nanopublications with CiTO citation intent annotations, than we can start formalizing intent annotations from a much wider range of use cases. For example, we can annotate historically important discussions. Anyone can retrospectively annotate all their own articles, making them more FAIR. And if we use DOI links, then it no longer is limited to journal articles, but we can use of for software and data citations too. This is where a recent template comes in created by Tobias Kuhn, one of the main nanopub developers:
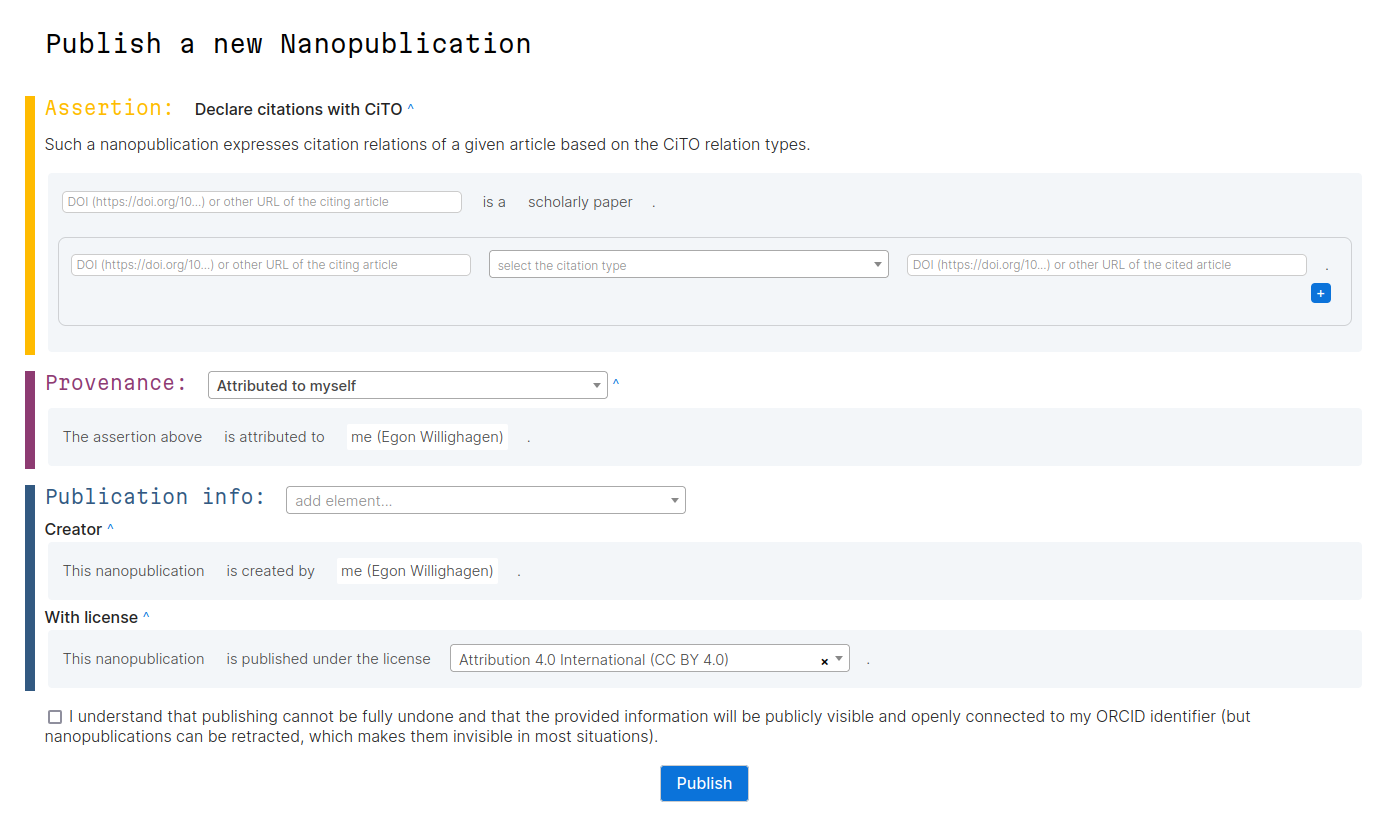
This nanopublication template defines the minimal needs of the assumptions, along with useful provenance and nanopub info. Basically, the assertion defines that one DOI is a ScholarlyWork and using the CiTO, defines that it cites one or more article works (with DOI). For each citations, one can select any of the known CiTO intent types, e.g. ‘extends’ or ‘uses method’ in, as in this nanopublication created with this template:
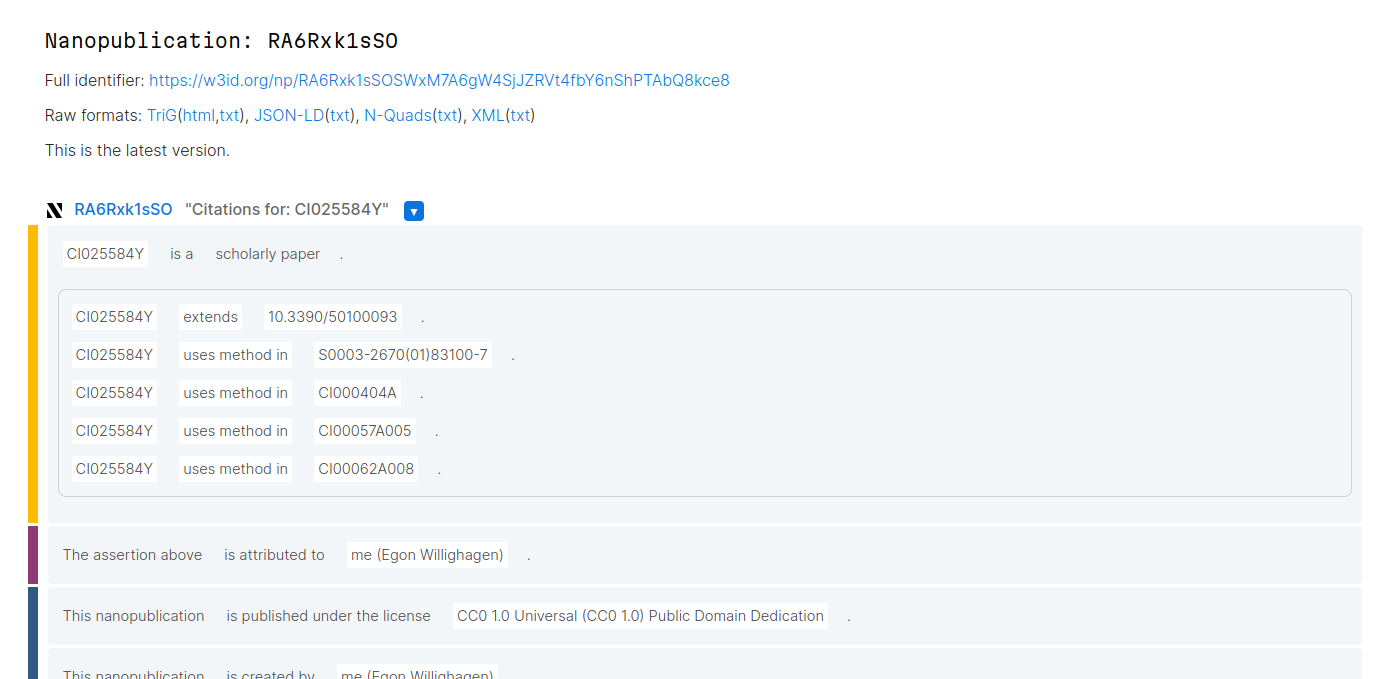
SPARQL-ing CiTO annotations
Besides the template, Tobias also started a SPARQL query to which I added restrictions that the citing and cited resources needs to have a DOI, giving us this query:
prefix rdfs: <http://www.w3.org/2000/01/rdf-schema#>
prefix np: <http://www.nanopub.org/nschema#>
prefix npa: <http://purl.org/nanopub/admin/>
prefix npx: <http://purl.org/nanopub/x/>
prefix xsd: <http://www.w3.org/2001/XMLSchema#>
prefix dct: <http://purl.org/dc/terms/>
select ?np ?label ?subj ?citationrel ?obj ?date where {
graph npa:graph {
?np npa:hasValidSignatureForPublicKey ?pubkey .
?np dct:created ?date .
?np np:hasAssertion ?assertion .
optional { ?np rdfs:label ?label . }
filter not exists { ?npx npx:invalidates ?np ; npa:hasValidSignatureForPublicKey ?pubkey . }
filter not exists { ?np npx:hasNanopubType npx:ExampleNanopub . }
}
graph ?assertion {
?subj ?citationrel ?obj .
filter(regex(str(?citationrel), "^http://purl.org/spar/cito/.*$"))
filter(regex(str(?subj), "doi.org/10"))
filter(regex(str(?obj), "doi.org/10"))
}
}
This includes 6 citation intentions defined by 4 nanopublications added during the Open Science Retreat:
- RAUjZE1JMu by me for a paper by Marija Purgar
- RAXgI–5gc by Christian Meesters
- RATZNhd3l_j by Taichi Oichi
- RA6Q6wxSYy by Niklas Hohmann
From nanopublications to Wikidata
Now, this query also provides me with enough information to propagate the citation intent (a fact?) to Wikidata and cite the original nanopublication as reference. With a variation of the above SPARQL query, I can get the five most recent new nanopublications, convert them to QuickStatements, and then enjoy them in Wikidata. This is written up in this Bacting script.
The script needs to handle some situations. For example, it will not add items for DOIs not already in Wikidata. So, if neither of the two DOIs are known in Wikidata, then nothing gets added. If they both are, then it will add the citation intent. There are alternative solutions, but in practice that doesn’t matter and the QuickStatements is in all situations the same, and QuickStatements will only add the new information.
This is what it will look like in Wikidata:
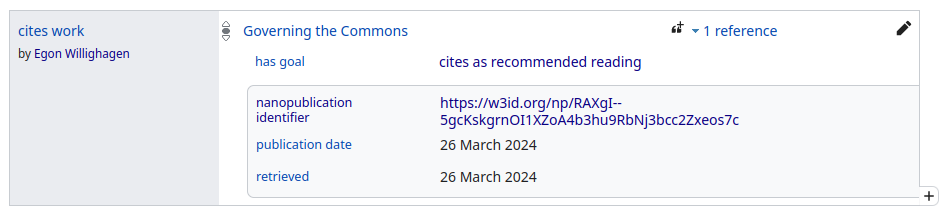
And this is what it looks (yellow) when we compare the contributions from nanopublications now with the other sources: