Oscar3, however, is capable of more then the name2structure of OPSIN (see also 10.1039/b411033a; it can take a plain text file with an experimental section with details on the synthesis of small organic compounds, and analyze the chemistry in that. This functionality has been available as an RSC authoring tool for some time now (see also 10.1039/b411699m). Unfortunately, what publisher put online (PDF and HTML) is much more difficult to process with Oscar3: those formats are often optimized for display, not for machine processing. The HTML can be cleaned up, but there is no general approach.
Christoph Steinbeck is going to present at the upcoming ACS meeting the use of Oscar3 for extraction of NMR spectra from old journal article, in preperation for submission to the NMRShiftDB.org (see the abstract of CINF 101).
Since the full Oscar3 was not hooked into Bioclipse yet, I had some work to do. It took me some time to figure out how to properly configure Oscar3, and what additional things I had to do to clean up the HTML used by publishers to get Oscar3 to extract NMR spectra (thanx to PeterC for hints!). I also had to tweak the Oscar3 code itself here and there, but that’s what opensource is about :) (Peter, if you are reading this: I have a number of patches for the Oscar3 code in bc_oscar; let me know if you’re interested in them.)
This is the end result:
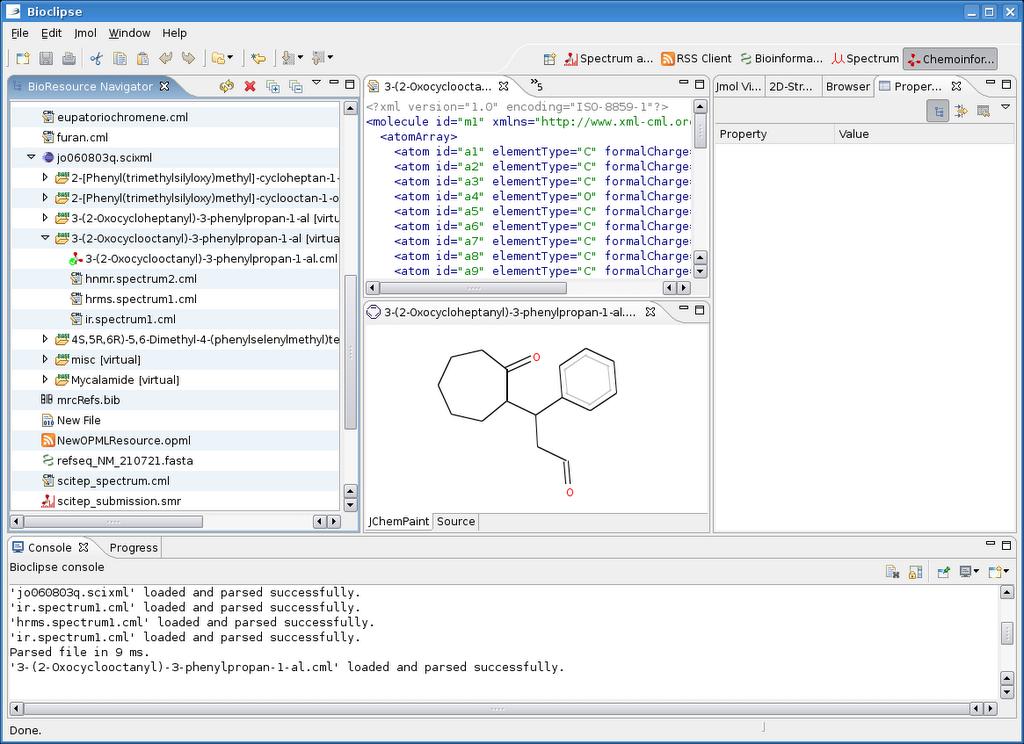
Note especially the hierarchy in the resource navigator on the left. The misc folder contains all the chemistry found in the article. But more importantly is that for six molecules it fully detected he experimental section! For 3-(2-Oxocyclooctanyl)-3-phenylpropan-1-al (InChI=1/C17H22O2/c18-13-12-15(14-8-4-3-5-9-14)16-10-6-1-2-7-11-17(16)19/h3-5,8-9,13,15-16H,1-2,6-7,10-12H2) it derived the molecular structure (with OPSIN), and a few spectra: H-NMR, high-resolution MS and IR.
So, if you attend the ACS meeting: make sure to visit Christoph’s CINF 101 presentation!
]]>