Practically, what matters here is how to ask the question and how to understand the answer.
We are supported in these practicalities with JavaScript libraries, in particular the ops.js library and general JSON functionality provided by most browsers (unless the student decided to use a different programming language, in which there are different libraries). Personally, I have only very limited JavaScript experience, and this mostly goes back to the good old Userscript and Greasemonkey days (wow! the paper is actually the 4th highest scoring BMC Bioinformatics article!). But because my JavaScript knowledge is limited and rusty, I spent a good part of today, to get a basic example running. Very basic, and barely exceeding the communication details. That is, this is the output in the browser:

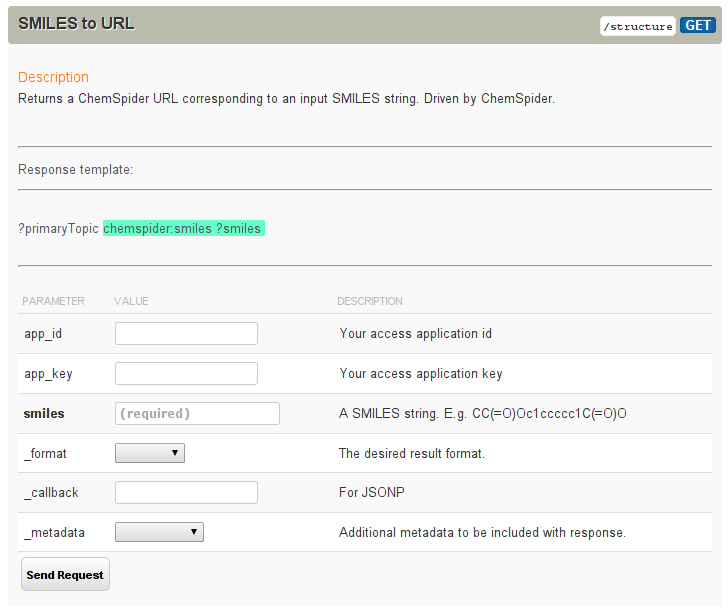
So, what does the question look like? The question is actually hardcoded in the HTML source, but the page does take two parameters:
the app_key and app_id that come with your Open PHACTS account.
The ops.js library helps us, and wraps the Open PHACTS LDA methods in JavaScript methods. Thus, rather can crafting special HTTP calls, we use two JavaScript calls:
var searcher = new Openphacts.ConceptWikiSearch(
"https://beta.openphacts.org",
params["app_id"], params["app_key"]
);
searcher.byTag(
'Aspirin', '20', '4', '07a84994-e464-4bbf-812a-a4b96fa3d197',
callback
);
The first statement creates an LDA method object, while the second makes an actual question. I have not defined the callback variable, which actually is a JavaScript function that looks like:
var callback = function(success, status, response){
var result = searcher.parseResponse(response);
document.getElementById("output").innerHTML =
"Results: " + JSON.stringify(result);
};
When the LDA web service returns data, this method gets called, providing asynchronous functionality to keep the web page responsive. But when called, it first parses the returned data, and then puts the JSON output as text in the HTML. The output that is given in the earlier screenshot.
So, hurdle taken. From here on it’s easier. Regular looping over the results, creating some HTML output, etc. The full source code if this example is available as Gist.
]]>