Day 3
Kai worked hard on getting the cdk.interfaces API cleaned up, as agreed upon earlier.
Christian added a test for the RMSD calculator
(see getAllAtomRMSD()), and cleaned up his code a bit. Stefan continued his bug-squashing on JChemPaint and fixed another one or two bugs.
Rajarshi uploaded a patch to set undefined atomic properties, like partial and formal charges and the implicit hydrogen count, to UNSET by default.
However, this broke the CDK at many places, as apparently many class methods assume the default to be zero. After discussing the issue at the CUBIC,
it turned out that this was sort of the intended, though undocumented, behavior: use the default Java values.
And I added missing clone() methods, closing one bug on SourceForge, added files for Eclipse to know how to build the CDK with Ant (thanx
to Nico for similar files for Jmol), and got CDK compiled again against Classpath.
Day 4
Miguel uploaded his first patched for support saving PDBPolymer data structures into and restoring them again from CML, addressing an almost two-year-old bug. He created new cdk.interfaces for them, to address module dependencies, but a large set of JUnit tests are yet missing.
Kai continued his cdk.interfaces refactoring, working on the more involved changes. Stefan, Tobias, and me worked on a poster and three three-fold flyers for our CDK booth at CompLife2006, so have not been very productive in bug squashing. But we are happy with the result. Below is a screenshot on one side of the main CDK folder:
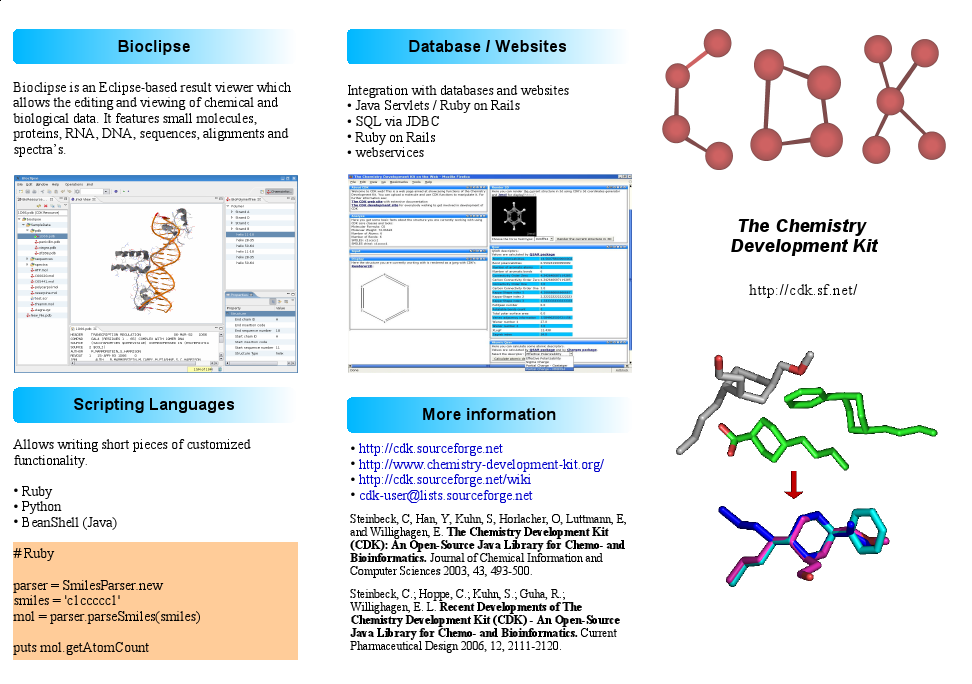
With 77 failing JUnit test, and still a too large number of open bugs on SourceForge, there is plenty of things to do today.
]]>