Then Google Scholar Citations (GSC) shows up. While it does not look as pretty as competing products, it compensates that with a wide coverage of literature (for example, it supports the JChemInf, which Web-of-Science currently does not; and I happen to publish a lot in that journal recently), books, and reports, while keeping false positives fairly low. Thus, it provides an upper limit of my citations statistics, but one I am pretty happy confident about. And my H-index is quite comparable anyway. This is what my profile looks like:
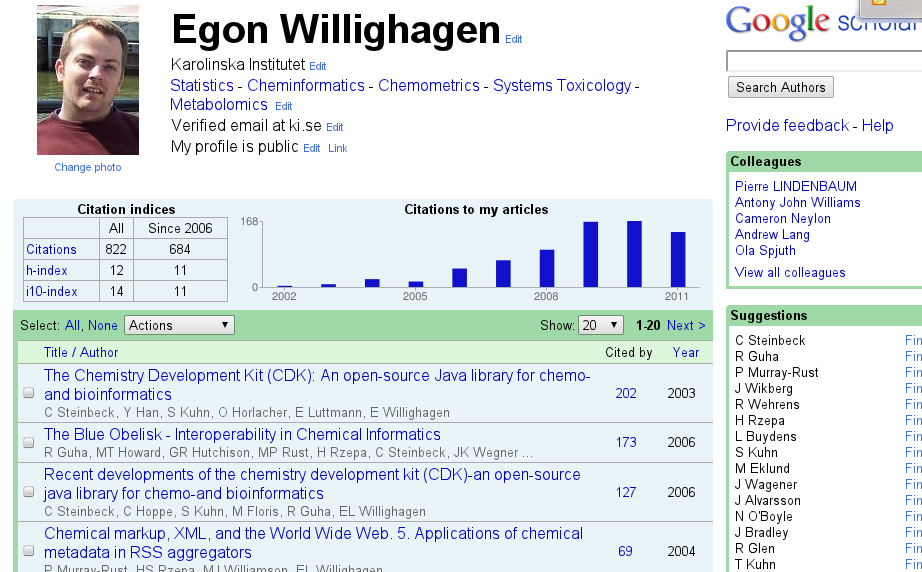
So, these statistics have two purposes to me: 1. grant applications, and 2. I like to know what people based theirs on my research. (Well, OK, 3. it helps me understand why I work so hard on too many things.)
Now the question is, will GSC take off. Will it replace ORCID? Will they join ORCID? Will GSC get a good API? Who will write the first userscript to make the GUI fancier? Will GSC support CiTO? Will GSC start using microformats or RDFa? What mashups can we expect between bibliographic databases? Will new entries automatically be posted to Google+? Will it have a button to autocreate a blog post when a paper gets cited 100, 500, or a 1000 times? Will GSC support #altmetrics?
]]>