In the afternoon I walked around a bit more in Oxford, did some more shopping… and visited the Apple shop and played with an iPad. It’s indeed a great piece of hardware. Looking forward to the first Android versions :)
The theme of the meeting was data, and this resulted in a wealth of presentations with cheminformatics. What is striking here is that a lot of work has not changed so much in 20 years, except for the scale. What I missed here was the large open data sets, but generally the level of open science was heartwarming! So many preprints mentions, GitHub repositories, and Zenodo deposits. The Blue Obelisk was truly ahead of its time, but it is a delight to see the field of chemistry catch up. I can now say a lot of about peer review, and why the field is not benefitting from all the experience that exists in the field because people publish in the wrong journals, but that is for another time.
I attended multiple sessions, which is a bit of a challenge, doing this remotely from Central European Summer Time (CEST).
Of course, the Sunday started with the Chemical informatics (R)evolution: Towards Democratization and Open Science
session, where I had my first talk, and later that day the Enhance your Data - Smart Ways to Metadata and Knowledge Graphs session,
where I gave a second talk, about Bioschemas’ ChemicalSubstance and MolecularEntity. Sadly, I
had to leave that meeting early because it was getting too late.
There were so many interesting sessions, I could not attend everything. I also have to go back to all my notes and isolate things I want to follow up on, prominently open datasets.
More later.
]]>In the afternoon I walked around a bit more in Oxford, did some more shopping… and visited the Apple shop and played with an iPad. It’s indeed a great piece of hardware. Looking forward to the first Android versions :)
Anyways, going to the Predictive Toxicology workshop, thanx to the bursary award I received from echeminfo (see Oxford, August 2010: eCheminfo Predictive ADME & Toxicology 2010 Workshop).
This afternoon I walked around a bit, watching all the old buildings. But I guess being here without anyone to share it with, and that it looks just like Cambridge, makes me not-so-much impressed. Moreover, it’s too busy with tourists and people randomly wearing Oxford University sweatshirts. Small and nice was the Museum of the History of Science, with some nice chemical pieces, like this one:
Buildings like the Radcliffe Camera are nice on the outside, but closed. Seems I have to become a fellow first. This is what it looked like today:
Quite interesting too was the Oxford University Press shop. I’m a sucker for books. Apparently, you can just write a book and publish it. For example, an extensive list of dictionaries on about anything… and since I have been writing several book chapters right now, perhaps this is actually an interesting route…
But the question is, of course, how long will we keep reading books… they’re the hamburgers of educational material… Kindle and alikes will soon drop in price, and cost some €30 euro. But e-book prices will have to drop too, and I still do not get why an e-book is more expensive than a paperback… (see Amazon, the Kindle edition is more expensive than the paperback??). But then again… they are rich, and I am not.
There was some recent talk about the fact that no one can be Open to the full. You either do Open Data or Open Source, and make a living from the rest. That’s where I nicely show I know bullocks of economics. I do BODR, CDK, … all Open, all for free.
OK. That’s a plus for Oxford… it makes you think about things. Perhaps there is something to morphogenetic fields…
]]>After the coffee break, Kuhn showed a coarse grained force field, approximating molecules by hacking them up in fragment of 3-10 heavy atoms. I guess, a bit like some small molecules force fields do for methyls. Fragments within a molecule are tied together by springs, and intra- and intermolecular force field parameters by running MD runs on fragment pairs. Varnek argued that QSPR for melting point prediction has reached a fundamental limited, with an RMSE of around 30 to 40 degrees Celsius, which makes it quite unreasonable to decide whether a compound with a predicted melting point of 40 degrees is solid or fluid at room temperature.
You have to forgive me for not reporting on the afternoon session; I was tied up talking with people at our booth, talking about the CDK, Taverna, Bioclipse, Jmol, other opensource chemoinformatics tools, and chemoinformatics in general. Very nice, but exhausting. I might advise the organization to set up a blog aggregator next year, though I am not sure whether there are others blogging about this conference.
]]>The conference is in honor of the life work by Prof. Gasteiger, who gave an overview of chemoinformatics in his group, Germany and Europe. He stressed the need of education in chemoinformatics, like in Obernai. He also highlighted that we, today, are still solving the same problem as 30 years ago. Which is true, which is why this channel is called Chem-bla-ics , trying to solve that problem. When asked if opensource chemoinformatics form the start would have addressed this, he replied that he requires people to cooperatively do research with his group; opensource clearly cannot enforce that.
Todays program had a number of interesting presentations (I, unfortunately, missed the first presentation, so have to visit that group soon now, to make up for that.) Prof. Aires-de-Sousa showed his work on MOLMAP for mapping metabolic networks (KEGG really, see my earlier blog ), and showed, just as proof of principle, classification of organisms based on this.
J. Weisser talked about docking, still an obligatory topic. This work really showed two new approaches: the use of QM partial charges (the example showed an improvement in RMSD of a factor 10, not very statistical, but promising indeed); the second was the fact that water does not like to be in tight spots, because of reduced possibilities for hydrogen bonding. A concept common in understand supramolecular phenomenon, but I have not seen this applied to docking before. But I am no expert in that field. M. Wagner showed work on using KEGG data to estimate likely metabolites, and the use in reducing effects of metabolic degradation. T. Schroeter introduced me to gaussian processes, a new data modeling method. Quite embarrassing to get introduced to such, as being specialized in modeling methods for chemical problems.
The poster session was, as normally, really exhausting, talking to a lot of people. Having a booth at the exhibition on opensource chemoinformatics added a nice twist to this. I therefore skipped the FIZ-award winner lectures, so I hope someone else will blog about those.
One last note: Sun started releasing their Java platform under the GPL license. Jim, seems that they proved me wrong . The class library is still not GPL, but is expected to become licensed such somewhere in the first half of next year.
]]>// to have short identifiers
Array = Packages.java.lang.reflect.Array;
String = Packages.java.lang.String;
msgBox = Packages.net.bioclipse.plugins.bc_rhino.ShowBcMsgBox;
DbfetchServiceServiceLocator =
Packages.uk.ac.ebi.www.ws.services.urn.Dbfetch.DbfetchServiceServiceLocator;
// get data
service = new DbfetchServiceServiceLocator();
strarray = service.getUrnDbfetch().fetchData("refseq:NM_210721", "refseq", "raw");
// make readable
str = new String();
for (i = 0; i < Array.getLength(strarray); i++) {
if (i != 0)
str = str + ("\n");
str = str + strarray[i];
}
// show
msgBox.ShowStatic(str);
It’s just a short example that uses webservice technology in Bioclipse to fetch a sequence.
QSAR support is getting along too, with a new DescriptorProvider extension point in trunk/ and work is progressing on a wizard that allows selecting descriptors and a CDK backend. The output of the wizard is a matrix resource, for which we already have a rich editor. A JOELib plugin has been suggested, as it has a good deal of QSAR descriptors too; Jörg, interested in doing a tiny bit of Bioclipse hacking?
A full proceedings is available online.
]]>About the last, the Renderer2D needs a serious overhaul. That is, a complete rewrite in proper Java2D, which can use affine transformations for zooming, scaling and fixing the coordinate system. The current code is ancient and predates Java2D. Rich’ code might be a good starting point. I would love to do this rewrite, but lack the resources… anyone in need of some open source fame?
I worked on atom typing, which is yet largely untested, and often integrated with other bits of code. Yesterday I uploaded some first patches which I wrote on the train ride back to the Netherlands.
Now, what can be concluded from this BSP? The participant count was below what I had hoped for, but those who did worked hard (and with pleasure I hope :) The total number of JUnit test has increased:
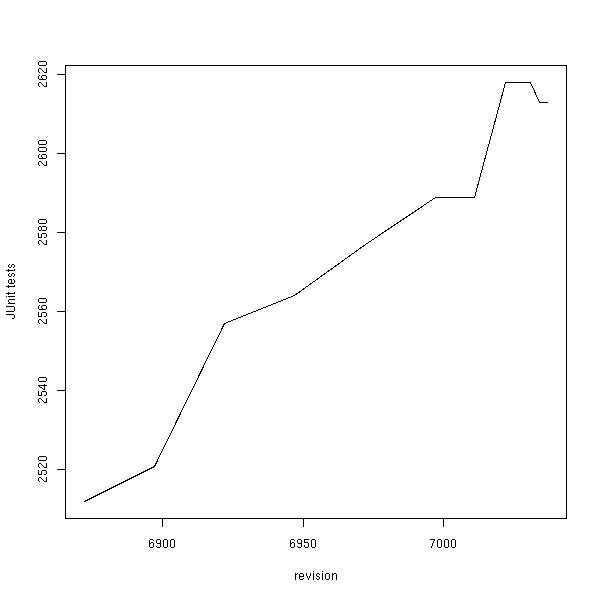
And so has the number of failing tests:
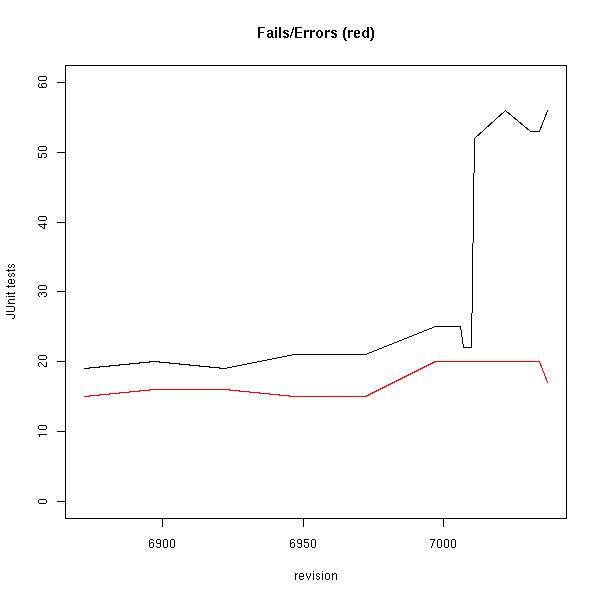
These plots were made with R from data created with two custom scripts both found in cdk/tools: makeBugCountPlot.pl and extractBugCountPlotData.bsh. Note that 96.86% of the tests do not fail!
The bump in failing tests seems to be due to commit 7010-7011, which has to do with SMILES parsing. Yes, the bond order resolving is still not solved. I don’t seem to get Todd’s patch for this working, but not giving up either. The bump is so large, because quite some JUnit tests use the SmilesParser as a quick tool to get a configured connection table. However, these tests should be replaced by explicit CDK models, which is easy done with the CDKSourceCodeWriter. I’ll blog about how to use that soon.
]]>Kai worked hard on getting the cdk.interfaces API cleaned up, as agreed upon earlier.
Christian added a test for the RMSD calculator
(see getAllAtomRMSD()), and cleaned up his code a bit. Stefan continued his bug-squashing on JChemPaint and fixed another one or two bugs.
Rajarshi uploaded a patch to set undefined atomic properties, like partial and formal charges and the implicit hydrogen count, to UNSET by default.
However, this broke the CDK at many places, as apparently many class methods assume the default to be zero. After discussing the issue at the CUBIC,
it turned out that this was sort of the intended, though undocumented, behavior: use the default Java values.
And I added missing clone() methods, closing one bug on SourceForge, added files for Eclipse to know how to build the CDK with Ant (thanx
to Nico for similar files for Jmol), and got CDK compiled again against Classpath.
Miguel uploaded his first patched for support saving PDBPolymer data structures into and restoring them again from CML, addressing an almost two-year-old bug. He created new cdk.interfaces for them, to address module dependencies, but a large set of JUnit tests are yet missing.
Kai continued his cdk.interfaces refactoring, working on the more involved changes. Stefan, Tobias, and me worked on a poster and three three-fold flyers for our CDK booth at CompLife2006, so have not been very productive in bug squashing. But we are happy with the result. Below is a screenshot on one side of the main CDK folder:
With 77 failing JUnit test, and still a too large number of open bugs on SourceForge, there is plenty of things to do today.
]]>More interestingly, discussion on the developers mailing list on the patch by Todd Martin of the EPA to address deducing bond orders in SMILES parsing (the major source of current open bugs!). A problem seems to be when his tool should be called in the SmilesParser class.
More details on the proceedings can be found on the BSP wiki page.
]]>