- provide a Java API for identifier mapping
- provide ID mappings (two flavors: with and without semantic meaning)
- provide services (R package, OpenAPI webservice)
- track the history of identifiers
The last one is more recent and two aspects are under development here: secondary identifiers and dead identifiers. More about that in some future post. About the first and the third I am also not going to tell much in this post. Just follow the above links.
I do want to say something in this post about the actually identifier mapping databases, in particular those we distribute as Apache Derby files, the storage format used by the Java libraries. These are the files you download if you want mapping databases for PathVisio (doi:10.1371/journal.pcbi.1004085). BridgeDb has mapping files for various things and some example databases the data it maps between:
- genes and proteins: Ensembl, UniProt, NCBI Gene
- metabolites; HMDB, ChEBI, LIPID MAPS, Wikidata, CAS
- publications: DOI, PubMed
- macromolecular complexes: Complex Portal, Wikidata
The BridgeDb API is agnostic to the things it can map identifiers for.
Downloading mapping files: BridgeDb has an BioSchemas-powered web page with an overview of the latest released mapping files. It looks like this:
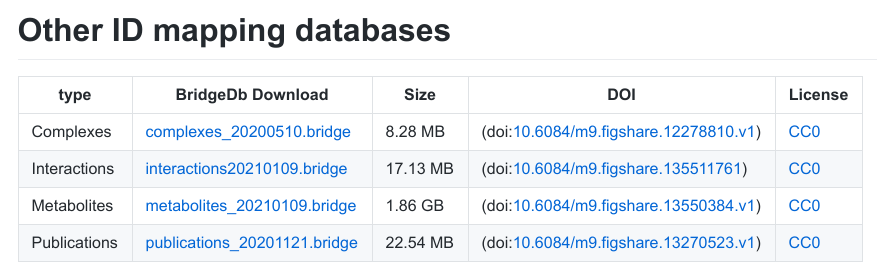
This webpage is the result from the cyber attack in late 2019, disrupting a good bit of the infrastructure. This is why we renewed the website, including the download page. The new page actually is hosted on GitHub as a Markdown file, but this is where things get interesting. The Markdown file is actually autogenerated from a JSON file with all the info. Everything, including the BioSchemas annotation is created from that. Basically, JSON gets converted into Markdown (with a custom script), which gets converted into HTML by a GitHub Action/Pages. So, when someone releases a new mapping file on Zenodo or Figshare, they only have to send me a pull request with updated JSON file.
Now, previously, downloading all released mapping files, for example for the BridgeDb webservice, was a bit complicated. The information was a HTML file generated by the webserver for a folder. No metadata. Nuno wrote code to extract the relevant info and download all the files. However, since the information is now available in a public JSON file, it is a lot easier. The following code uses wget and jq, two tools readily available on the popular operating systems. Have fun!
!/bin/bash
wget -nc https://bridgedb.github.io/data/gene.json
wget -nc https://bridgedb.github.io/data/corona.json
wget -nc https://bridgedb.github.io/data/other.json
jq -r '.mappingFiles | .[] | "\(.file)=\(.downloadURL)"' gene.json > files.txt
jq -r '.mappingFiles | .[] | "\(.file)=\(.downloadURL)"' corona.json >> files.txt
jq -r '.mappingFiles | .[] | "\(.file)=\(.downloadURL)"' other.json >> files.txt
for FILE in $(cat files.txt)
do
readarray -d = -t splitFILE<<< "$FILE"
echo ${splitFILE[0]}
wget -nc -O ${splitFILE[0]} ${splitFILE[1]}
done
Actually, while writing this blog post, I notice the code can be further simplified.
]]>