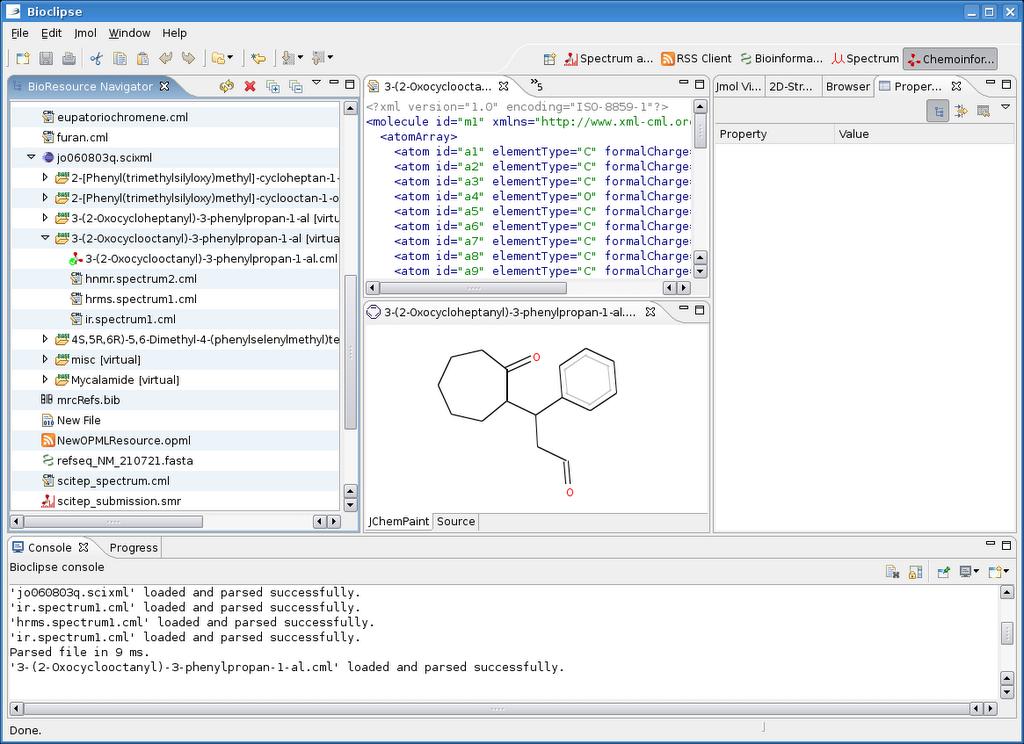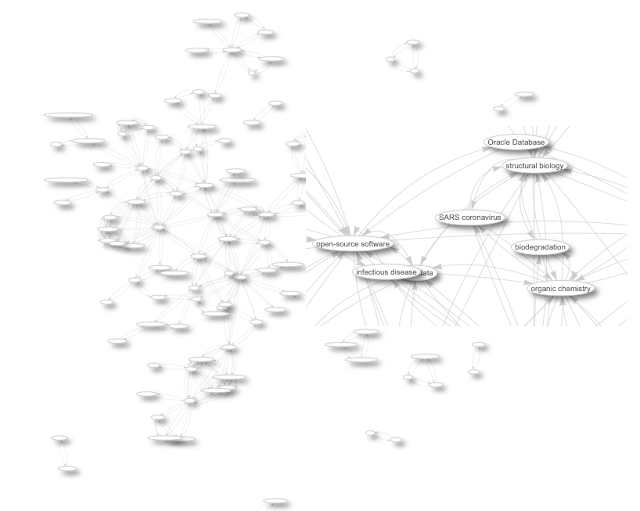
Graphs like this show information on how people are using my work, which in turn allows me to further support. But this relies on open citations.
In my opinion, citations are an essential part of our research process. It gives us access to import prior work on which a study is based, and reflects how a work influences other research or even is essential to that other work. For example, it allows us to not repeat earlier published work, while preserving the ability to reproduce the full work. The Initiative for Open Citations encourages these citations to be publicly available to benefit research, but removing barriers to access this critical part of scholarly communication. While many societies and publishers have joined this initiative, the American Chemical Society (ACS) has not yet. By not joining the limit the sharing of knowledge for unclear reasons.
And I would really like to see the ACS to join this initiative, and proposed this a few times already. Because they still have not joined the initiative, I have started this petition. If you agree, please sign and share it with others.
]]>